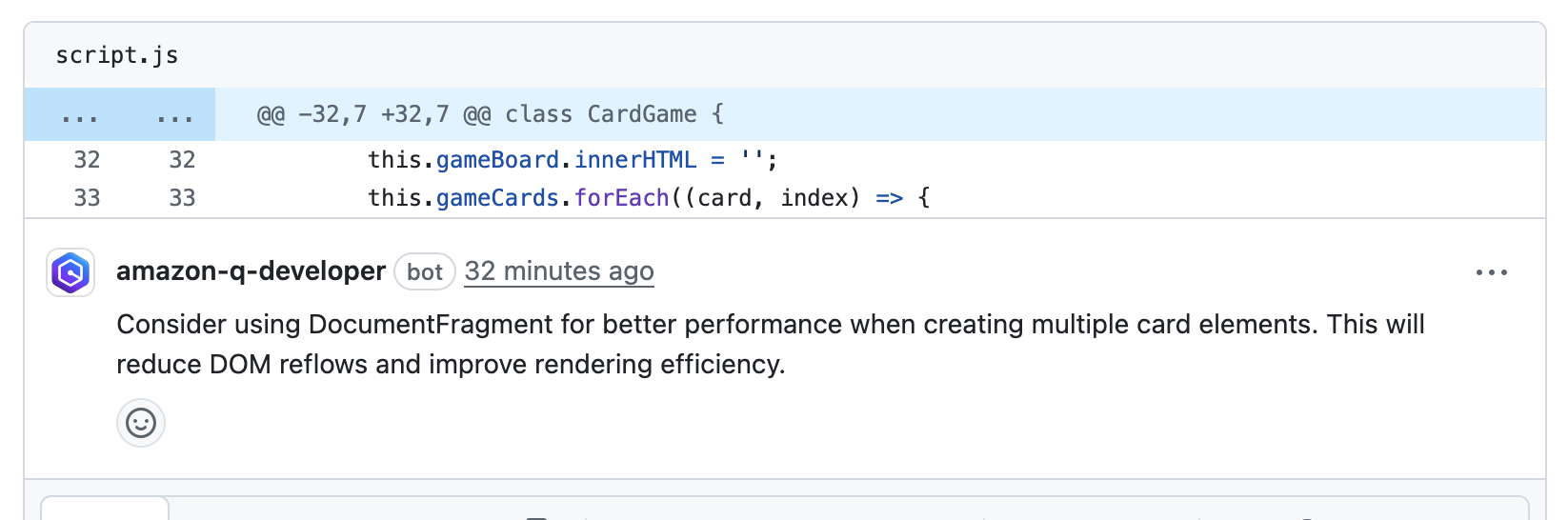
Post Syndicated from Akash Verma original https://aws.amazon.com/blogs/devops/automating-detection-of-security-vulnerabilities-and-bugs-in-ci-cd-pipelines-using-amazon-codeguru-reviewer-cli/
Watts S. Humphrey, the father of Software Quality, had famously quipped, “Every business is a software business”. Software is indeed integral to any industry. The engineers who create software are also responsible for making sure that the underlying code adheres to industry and organizational standards, are performant, and are absolved of any security vulnerabilities that could make them susceptible to attack.
Traditionally, security testing has been the forte of a specialized security testing team, who would conduct their tests toward the end of the Software Development lifecycle (SDLC). The adoption of DevSecOps practices meant that security became a shared responsibility between the development and security teams. Now, development teams can, on their own or as advised by their security team, setup and configure various code scanning tools to detect security vulnerabilities much earlier in the software delivery process (aka “Shift Left”). Meanwhile, the practice of Static code analysis and security application testing (SAST) has become a standard part of the SDLC. Furthermore, it’s imperative that the development teams expect SAST tools that are easy to set-up, seamlessly fit into their DevOps infrastructure, and can be configured without requiring assistance from security or DevOps experts.
In this post, we’ll demonstrate how you can leverage Amazon CodeGuru Reviewer Command Line Interface (CLI) to integrate CodeGuru Reviewer into your Jenkins Continuous Integration & Continuous Delivery (CI/CD) pipeline. Note that the solution isn’t limited to Jenkins, and it would be equally useful with any other build automation tool. Moreover, it can be integrated at any stage of your SDLC as part of the White-box testing. For example, you can integrate the CodeGuru Reviewer CLI as part of your software development process, as well as run it on your dev machine before committing the code.
Launched in 2020, CodeGuru Reviewer utilizes machine learning (ML) and automated reasoning to identify security vulnerabilities, inefficient uses of AWS APIs and SDKs, as well as other common coding errors. CodeGuru Reviewer employs a growing set of detectors for Java and Python to provide recommendations via the AWS Console. Customers that leverage the CodeGuru Reviewer CLI within a CI/CD pipeline also receive recommendations in a machine-readable JSON format, as well as HTML.
CodeGuru Reviewer offers native integration with Source Code Management (SCM) systems, such as GitHub, BitBucket, and AWS CodeCommit. However, it can be used with any SCM via its CLI. The CodeGuru Reviewer CLI is a shim layer on top of the AWS Command Line Interface (AWS CLI) that simplifies the interaction with the tool by handling the uploading of artifacts, triggering of the analysis, and fetching of the results, all in a single command.
Many customers, including Mastercard, are benefiting from this new CodeGuru Reviewer CLI.
“During one of our technical retrospectives, we noticed the need to integrate Amazon CodeGuru recommendations in our build pipelines hosted on Jenkins. Not all our developers can run or check CodeGuru recommendations through the AWS console. Incorporating CodeGuru CLI in our build pipelines acts as an important quality gate and ensures that our developers can immediately fix critical issues.”
Claudio Frattari, Lead DevOps at Mastercard
Solution overview
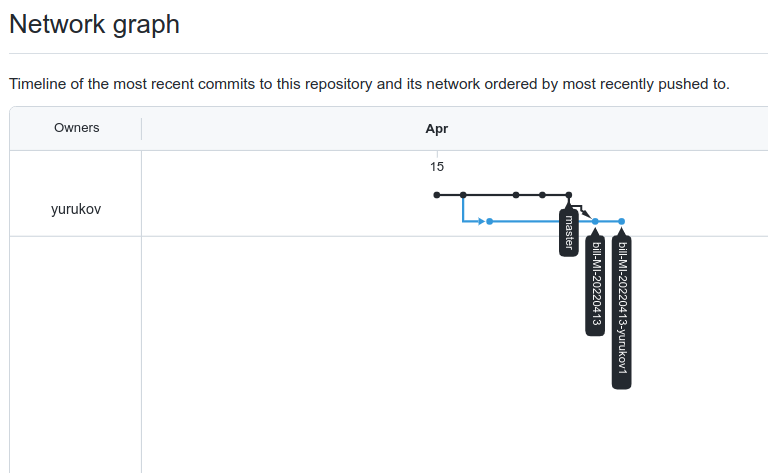
The application deployment workflow starts by placing the application code on a GitHub SCM. To automate the scenario, we have added GitHub to the Jenkins project under the “Source Code” section. We chose the GitHub option, which would clone the chosen GitHub repository in the Jenkins local workspace directory.
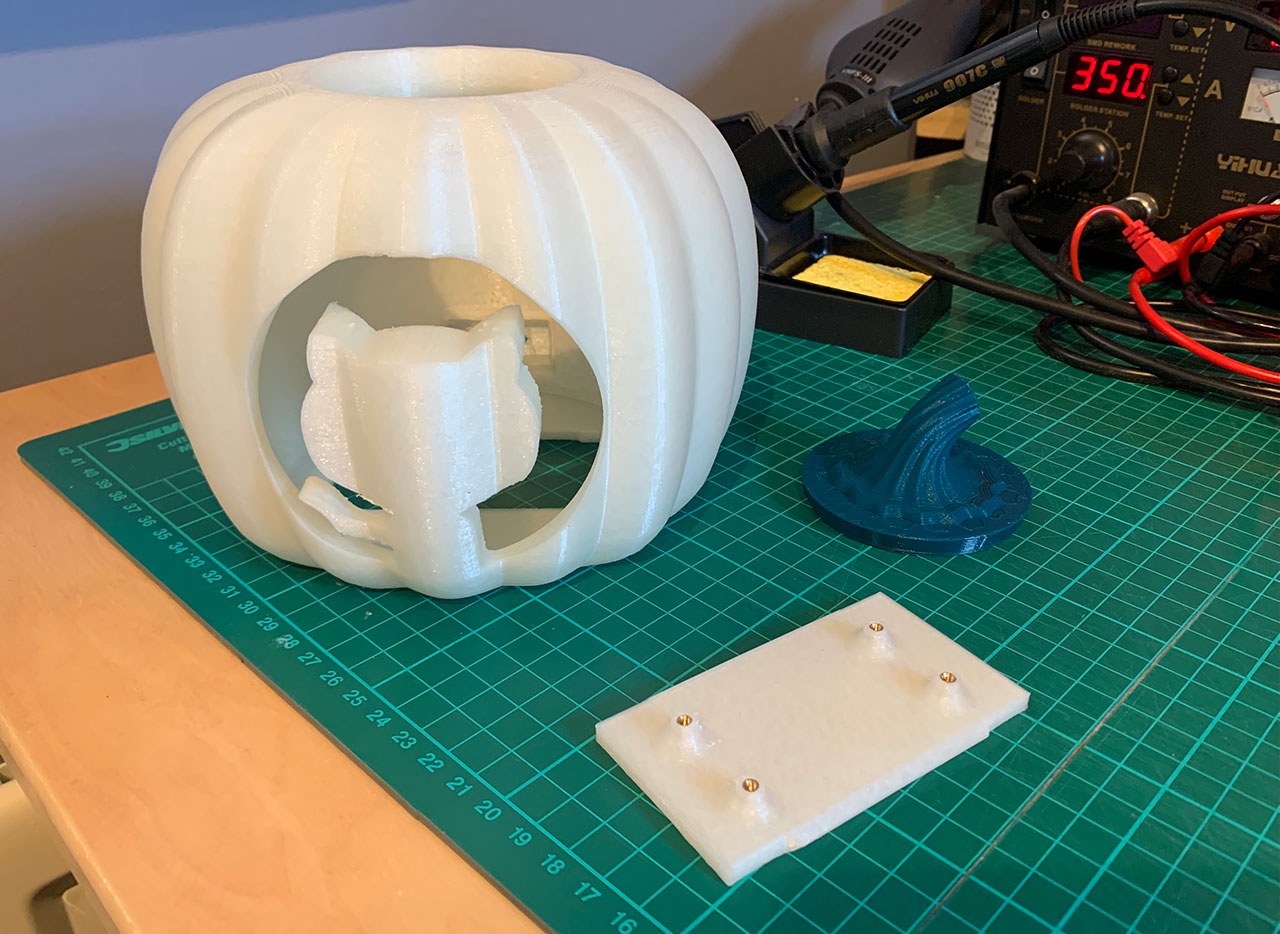
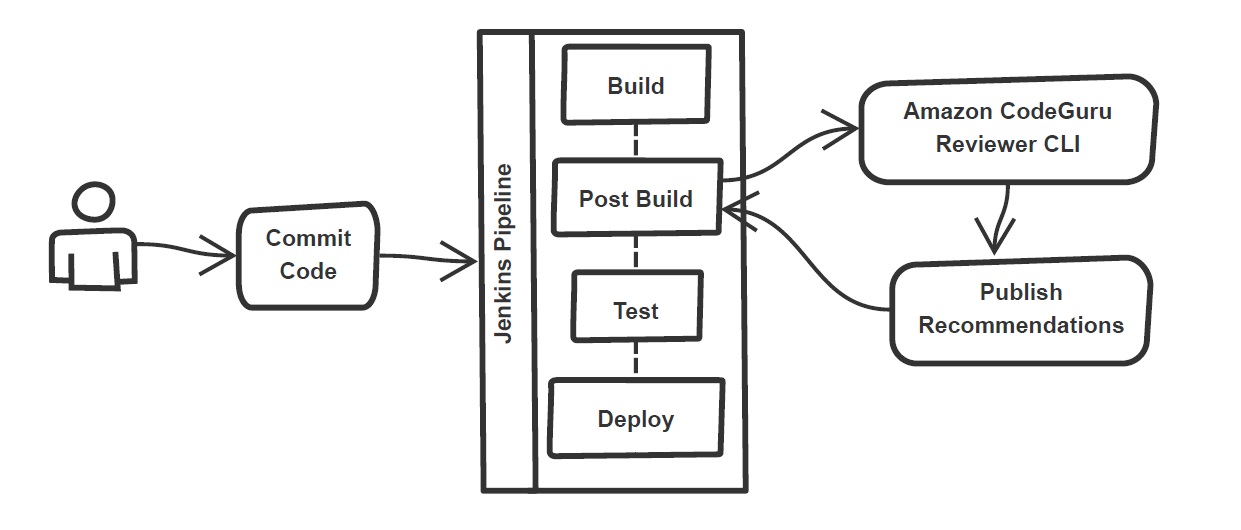
In the build stage of the pipeline (see Figure 1), we configure the appropriate build tool to perform the code build and security analysis. In this example, we will be using Maven as the build tool.

Figure 1: Jenkins pipeline with Amazon CodeGuru Reviewer
In the post-build stage, we configure the CodeGuru Reviewer CLI to generate the recommendations based on the review.
Lastly, in the concluding stage of the pipeline, we’ll be analyzing the JSON results using jq – a lightweight and flexible command-line JSON processor, and then failing the Jenkins job if we encounter observations that are of a “Critical” severity.
Jenkins will trigger the “CodeGuru Reviewer” (see Figure 1) based review process in the post-build stage, i.e., after the build finishes. Furthermore, you can configure other stages, such as automated testing or deployment, after this stage. Additionally, passing the location of the build artifacts to the CLI lets CodeGuru Reviewer perform a more in-depth security analysis. Build artifacts are either directories containing jar files (e.g., build/lib for Gradle or /target for Maven) or directories containing class hierarchies (e.g., build/classes/java/main for Gradle).
Walkthrough
Now that we have an overview of the workflow, let’s dive deep and walk you through the following steps in detail:
- Installing the CodeGuru Reviewer CLI
- Creating a Jenkins pipeline job
- Reviewing the CodeGuru Reviewer recommendations
- Configuring CodeGuru Reviewer CLI’s additional options
1. Installing the CodeGuru CLI Wrapper
a. Prerequisites
To run the CLI, we must have Git, Java, Maven, and the AWS CLI installed. Verify that they’re installed on our machine by running the following commands:
java -version
mvn --version
aws --version
git –-version
If they aren’t installed, then download and install Java here (Amazon Corretto is a no-cost, multiplatform, production-ready distribution of the Open Java Development Kit), Maven from here, and Git from here. Instructions for installing AWS CLI are available here.
We would need to create an Amazon Simple Storage Service (Amazon S3) bucket with the prefix codeguru-reviewer-. Note that the bucket name must begin with the mentioned prefix, since we have used the name pattern in the following AWS Identity and Access Management (IAM) permissions, and CodeGuru Reviewer expects buckets to begin with this prefix. Refer to the following section 4(a) “Specifying S3 bucket name” for more details.
Furthermore, we’ll need working credentials on our machine to interact with our AWS account. Learn more about setting up credentials for AWS here. You can find the minimal permissions to run the CodeGuru Reviewer CLI as follows.
b. Required Permissions
To use the CodeGuru Reviewer CLI, we need at least the following AWS IAM permissions, attached to an AWS IAM User or an AWS IAM role:
{
"Version": "2012-10-17",
"Statement": [
{
"Action": [
"codeguru-reviewer:ListRepositoryAssociations",
"codeguru-reviewer:AssociateRepository",
"codeguru-reviewer:DescribeRepositoryAssociation",
"codeguru-reviewer:CreateCodeReview",
"codeguru-reviewer:DescribeCodeReview",
"codeguru-reviewer:ListRecommendations",
"iam:CreateServiceLinkedRole"
],
"Resource": "*",
"Effect": "Allow"
},
{
"Action": [
"s3:CreateBucket",
"s3:GetBucket*",
"s3:List*",
"s3:GetObject",
"s3:PutObject",
"s3:DeleteObject"
],
"Resource": [
"arn:aws:s3:::codeguru-reviewer-*",
"arn:aws:s3:::codeguru-reviewer-*/*"
],
"Effect": "Allow"
}
]
}
c. CLI installation
Please download the latest version of the CodeGuru Reviewer CLI available at GitHub. Then, run the following commands in sequence:
curl -OL https://github.com/aws/aws-codeguru-cli/releases/download/0.0.1/aws-codeguru-cli.zip
unzip aws-codeguru-cli.zip
export PATH=$PATH:./aws-codeguru-cli/bin
d. Using the CLI
The CodeGuru Reviewer CLI only has one required parameter –root-dir (or just -r) to specify to the local directory that should be analyzed. Furthermore, the –src option can be used to specify one or more files in this directory that contain the source code that should be analyzed. In turn, for Java applications, the –build option can be used to specify one or more build directories.
For a demonstration, we’ll analyze the demo application. This will make sure that we’re all set for when we leverage the CLI in Jenkins. To proceed, first we download and install the sample application, as follows:
git clone https://github.com/aws-samples/amazon-codeguru-reviewer-sample-app
cd amazon-codeguru-reviewer-sample-app
mvn clean compile
Now that we have built our demo application, we can use the aws-codeguru-cli CLI command that we added to the path to trigger the code scan:
aws-codeguru-cli --root-dir ./ --build target/classes --src src --output ./output
For additional assistance on the CLI command, reference the readme here.
2. Creating a Jenkins Pipeline job
CodeGuru Reviewer can be integrated in a Jenkins Pipeline as well as a Freestyle project. In this example, we’re leveraging a Pipeline.
a. Pipeline Job Configuration
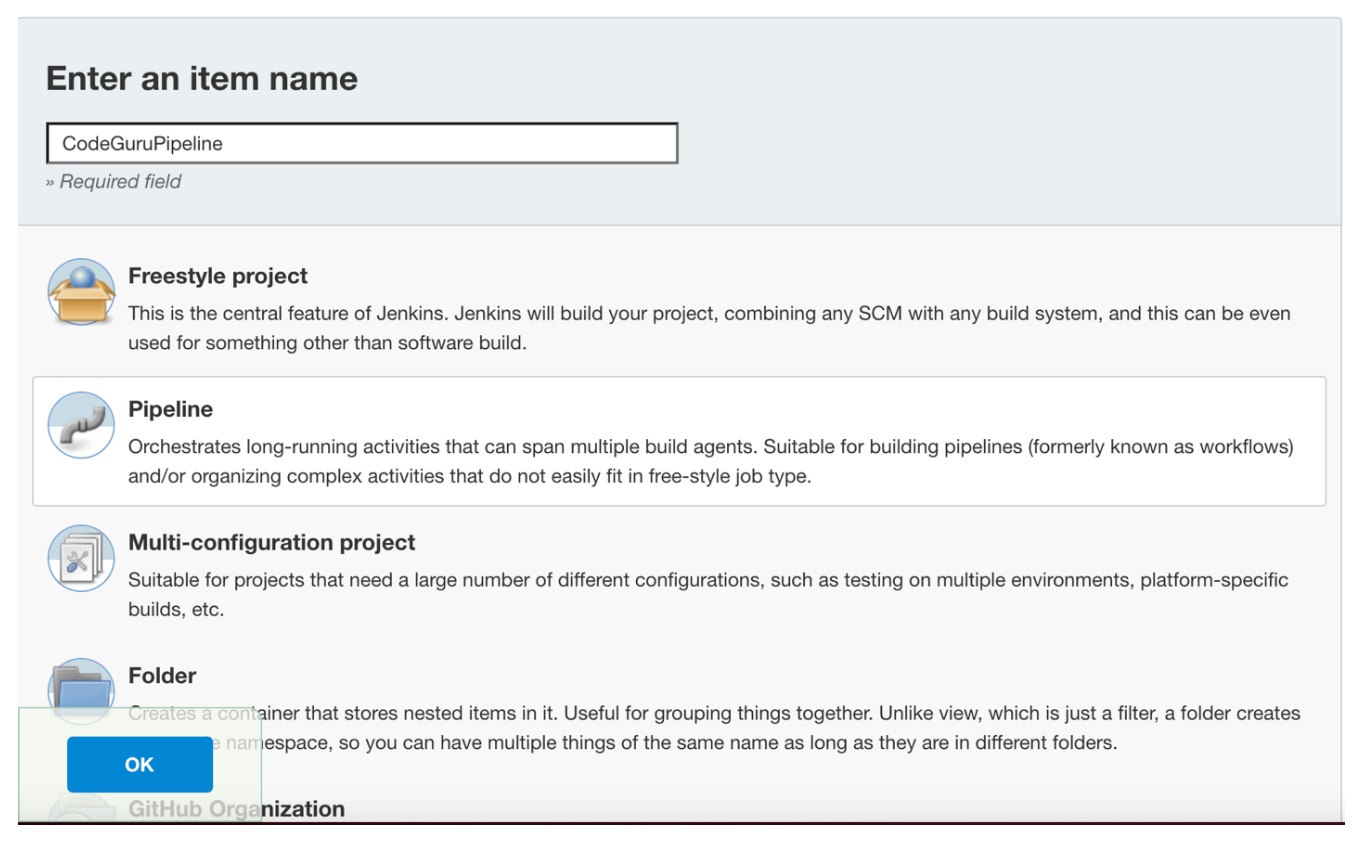
- Log in to Jenkins, choose “New Item”, then select “Pipeline” option.
- Enter a name for the project (for example, “CodeGuruPipeline”), and choose OK.

Figure 2: Creating a new Jenkins pipeline
- On the “Project configuration” page, scroll down to the bottom and find your pipeline. In the pipeline script, paste the following script (or use your own Jenkinsfile). The following example is a valid Jenkinsfile to integrate CodeGuru Reviewer with a project built using Maven.
pipeline {
agent any
stages {
stage('Build') {
steps {
// Get code from a GitHub repository
git clone https://github.com/aws-samples/amazon-codeguru-reviewer-java-detectors.git
// Run Maven on a Unix agent
sh "mvn clean compile"
// To run Maven on a Windows agent, use following
// bat "mvn -Dmaven.test.failure.ignore=true clean package"
}
}
stage('CodeGuru Reviewer') {
steps{
sh 'ls -lsa *'
sh 'pwd'
// Here we’re setting an absolute path, but we can
// also use JENKINS environment variables
sh '''
export BASE=/var/jenkins_home/workspace/CodeGuruPipeline/amazon-codeguru-reviewer-java-detectors
export SRC=${BASE}/src
export OUTPUT = ./output
/home/codeguru/aws-codeguru-cli/bin/aws-codeguru-cli --root-dir $BASE --build $BASE/target/classes --src $SRC --output $OUTPUT -c $GIT_PREVIOUS_COMMIT:$GIT_COMMIT --no-prompt
'''
}
}
stage('Checking findings'){
steps{
// In this example we are stopping our pipline on
// detecting Critical findings. We are using jq
// to count occurrences of Critical severity
sh '''
CNT = $(cat ./output/recommendations.json |jq '.[] | select(.severity=="Critical")|.severity' | wc -l)'
if (( $CNT > 0 )); then
echo "Critical findings discovered. Failing."
exit 1
fi
'''
}
}
}
}
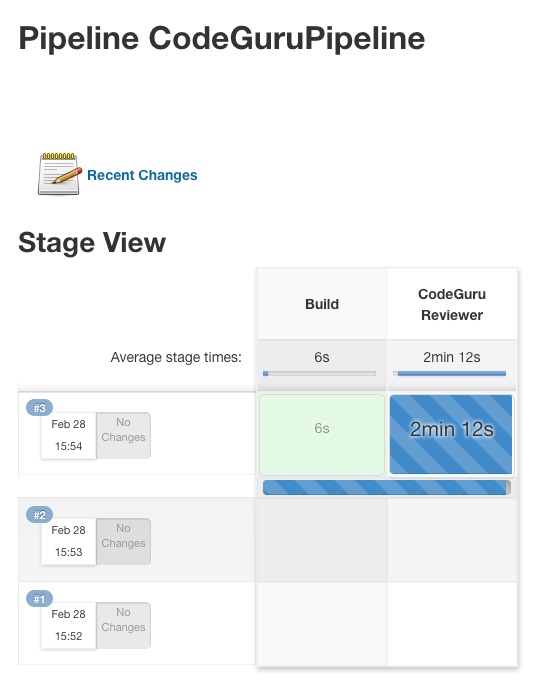
- Save the configuration and select “Build now” on the side bar to trigger the build process (see Figure 3).

Figure 3: Jenkins pipeline in triggered state
3. Reviewing the CodeGuru Reviewer recommendations
Once the build process is finished, you can view the review results from CodeGuru Reviewer by selecting the Jenkins build history for the most recent build job. Then, browse to Workspace output. The output is available in JSON and HTML formats (Figure 4).

Figure 4: CodeGuru CLI Output
Snippets from the HTML and JSON reports are displayed in Figure 5 and 6 respectively.
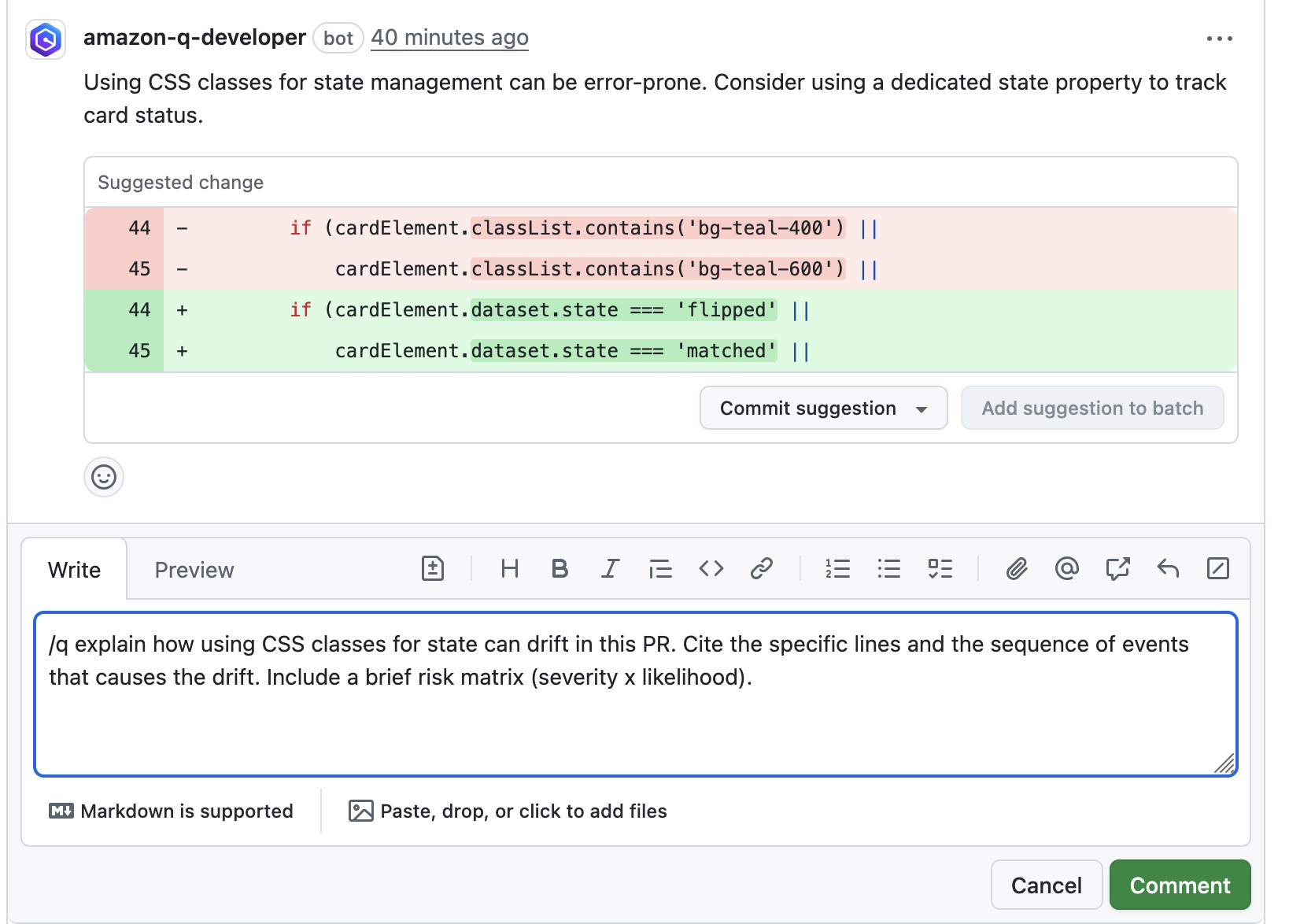
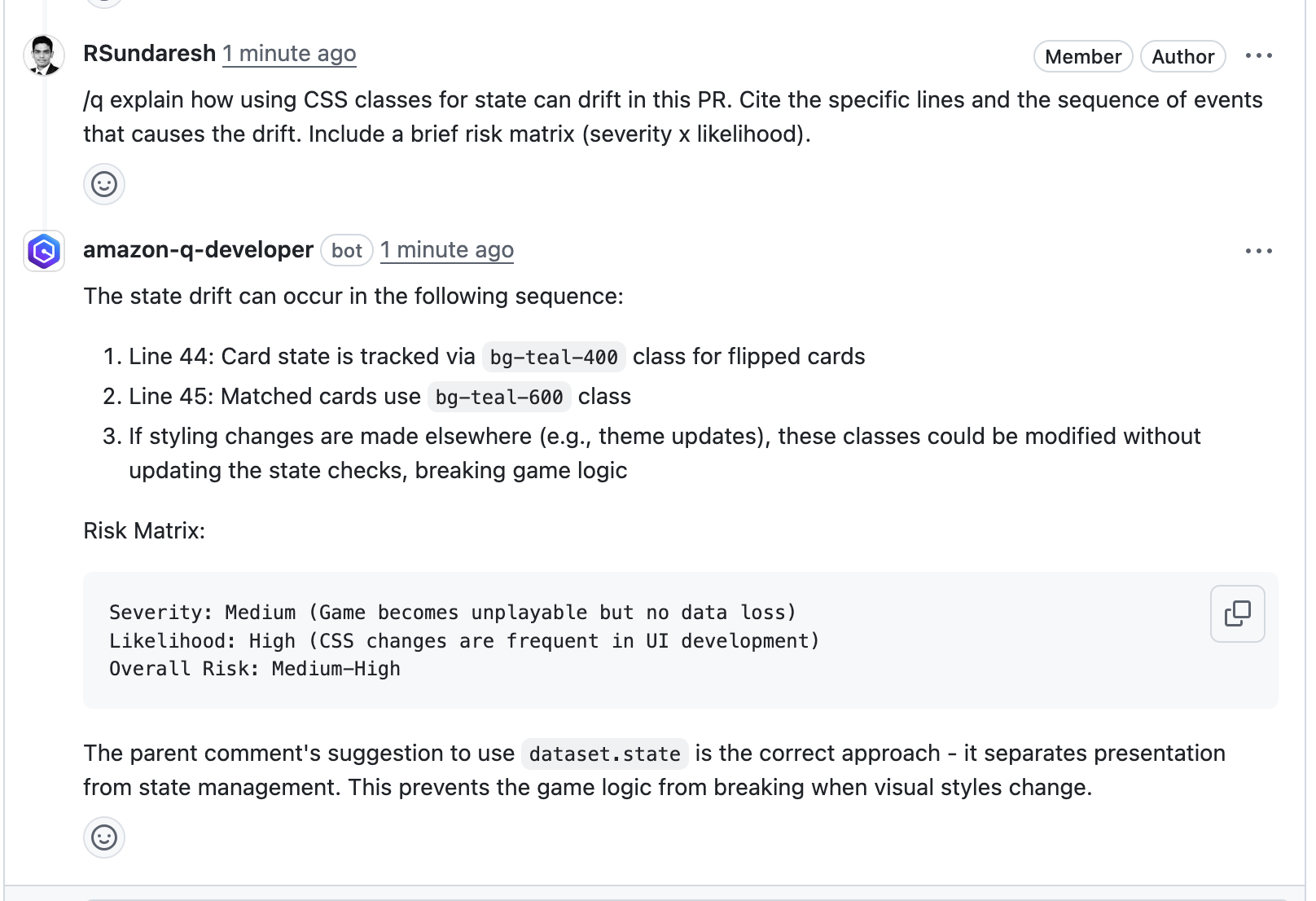
In this example, our pipeline analyzes the JSON results with jq based on severity equal to critical and failing the job if there are any critical findings. Note that this output path is set with the –output option. For instance, the pipeline will fail on noticing the “critical” finding at Line 67 of the EventHandler.java class (Figure 5), flagged due to use of an insecure code. Till the time the code is remediated, the pipeline would prevent the code deployment. The vulnerability could have gone to production undetected, in absence of the tool.

Figure 5: CodeGuru HTML Report

Figure 6: CodeGuru JSON recommendations
4. Configuring CodeGuru Reviewer CLI’s additional options
a. Specifying Amazon S3 bucket name and policy
CodeGuru Reviewer needs one Amazon S3 bucket for the CLI to store the artifacts while the analysis is running. The artifacts are deleted after the analysis is completed. The same bucket will be reused for all the repositories that are analyzed in the same account and region (unless specified otherwise by the user). Note that CodeGuru Reviewer expects the S3 bucket name to begin with codeguru-reviewer-. At this time, you can’t use a different naming pattern. However, if you want to use a different bucket name, then you can use the –bucket-name option.
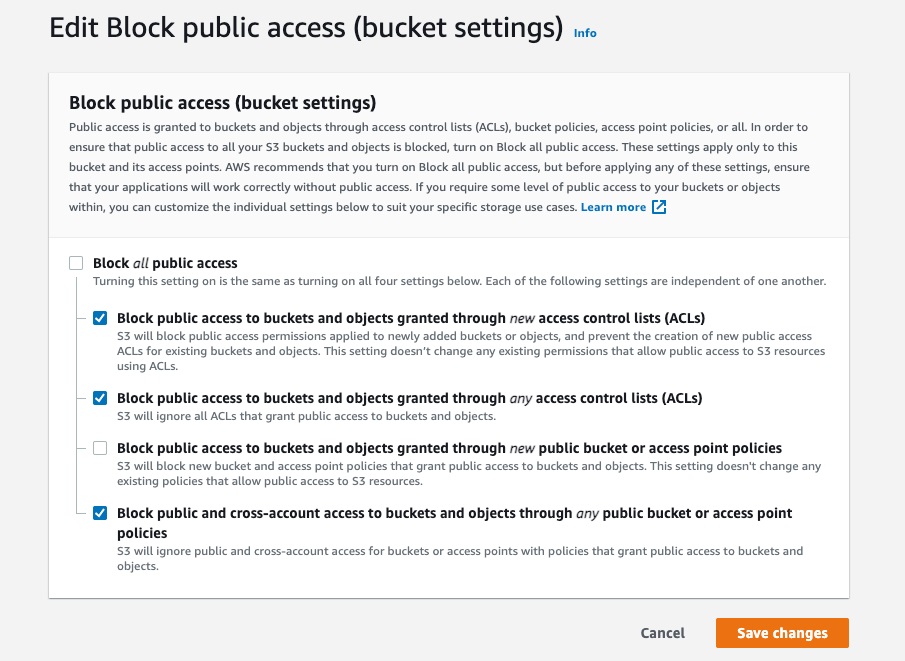
Select the Permissions tab of your S3 bucket. Update the Block public access and add the following S3 bucket policy.

Figure 7: S3 bucket settings
S3 bucket policy:
{
"Version":"2012-10-17",
"Statement":[
{
"Sid":"PublicRead",
"Effect":"Allow",
"Principal":"*",
"Action":"s3:GetObject",
"Resource":"[Change to ARN for your S3 bucket]/*"
}
]
}
Note that if you must change the bucket’s name, then you can remove the associated S3 bucket in the AWS console under CodeGuru → CI workflows and select Disassociate Workflow.
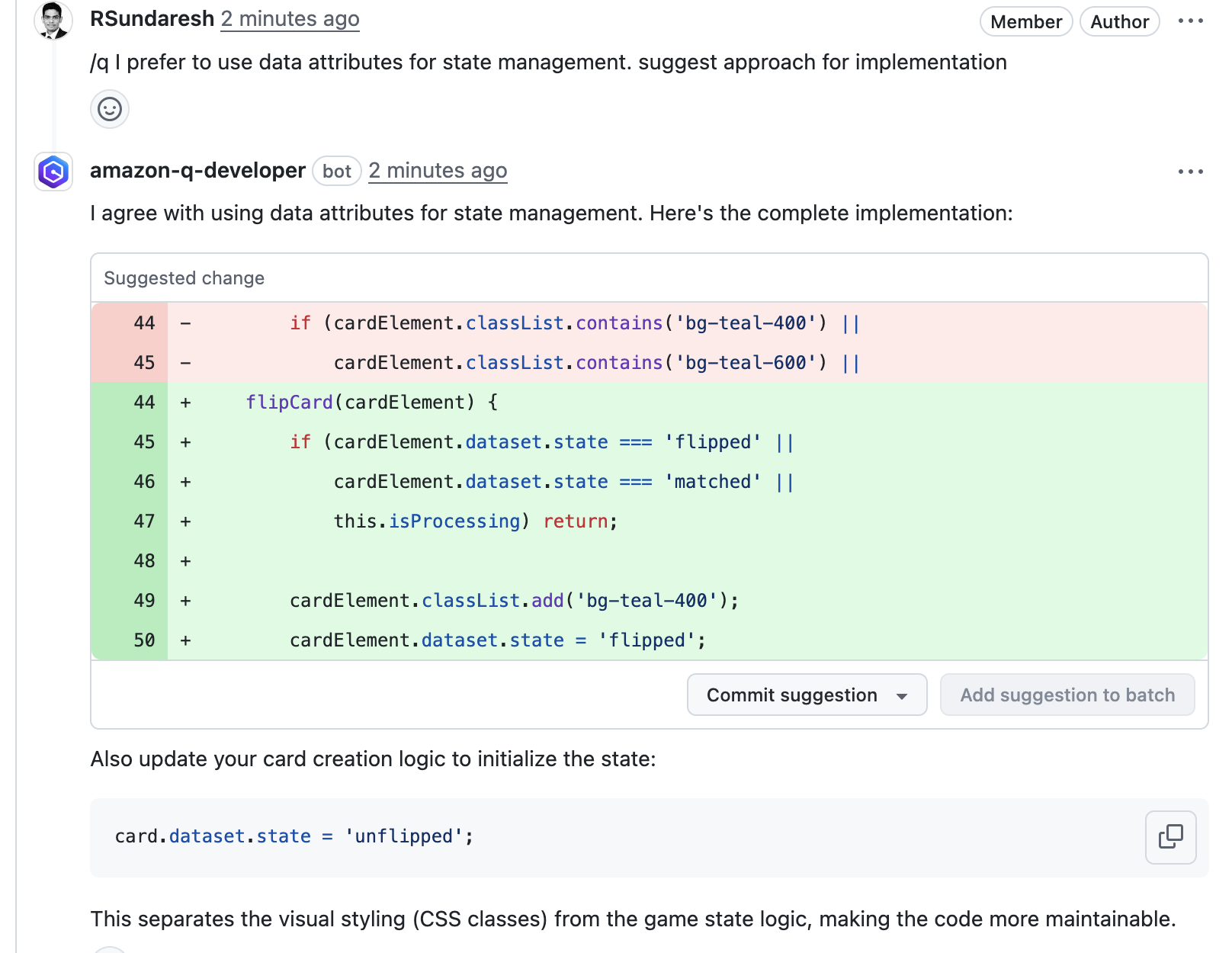
b. Analyzing a single commit
The CLI also lets us specify a specific commit range to analyze. This can lead to faster and more cost-effective scans for the incremental code changes, instead of a full repository scan. For example, if we just want to analyze the last commit, we can run:
aws-codeguru-cli -r ./ -s src/main/java -b build/libs -c HEAD^:HEAD --no-prompt
Here, we use the -c option to specify that we only want to analyze the commits between HEAD^ (the previous commit) and HEAD (the current commit). Moreover, we add the –no-prompt option to automatically answer questions by the CLI with yes. This option is useful if we plan to use the CLI in an automated way, such as in our CI/CD workflow.
c. Encrypting artifacts
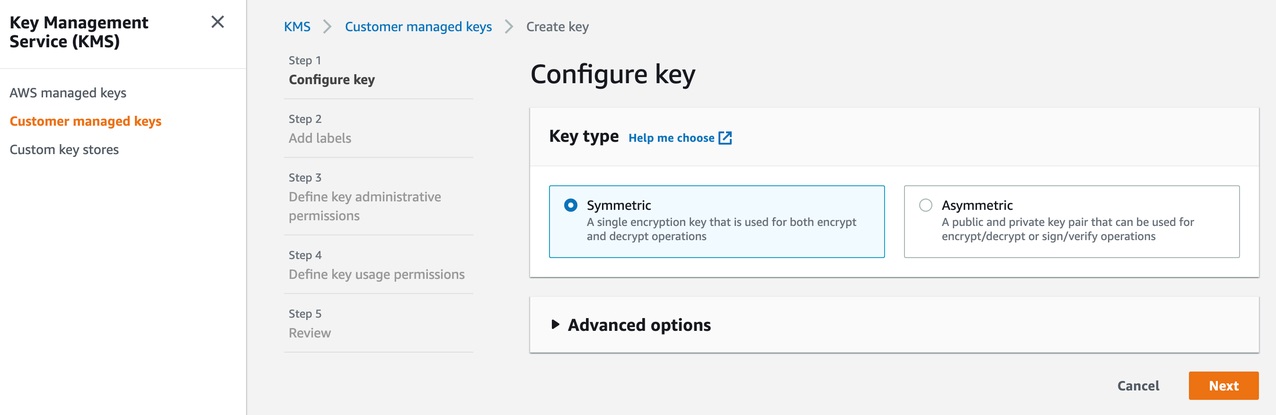
CodeGuru Reviewer lets us use a customer managed key to encrypt the content of the S3 bucket that is used to store the source and build artifacts. To achieve this, create a customer owned key in AWS Key Management Service (AWS KMS) (see Figure 8).

Figure 8: KMS settings
We must grant CodeGuru Reviewer the permission to decrypt artifacts with this key by adding the following Statement to your Key policy:
{
"Sid":"Allow CodeGuru to use the key to decrypt artifact",
"Effect":"Allow",
"Principal":{
"AWS":"*"
},
"Action":[
"kms:Decrypt",
"kms:DescribeKey"
],
"Resource":"*",
"Condition":{
"StringEquals":{
"kms:ViaService":"codeguru-reviewer.amazonaws.com",
"kms:CallerAccount":[
"YOUR AWS ACCOUNT ID"
]
}
}
}
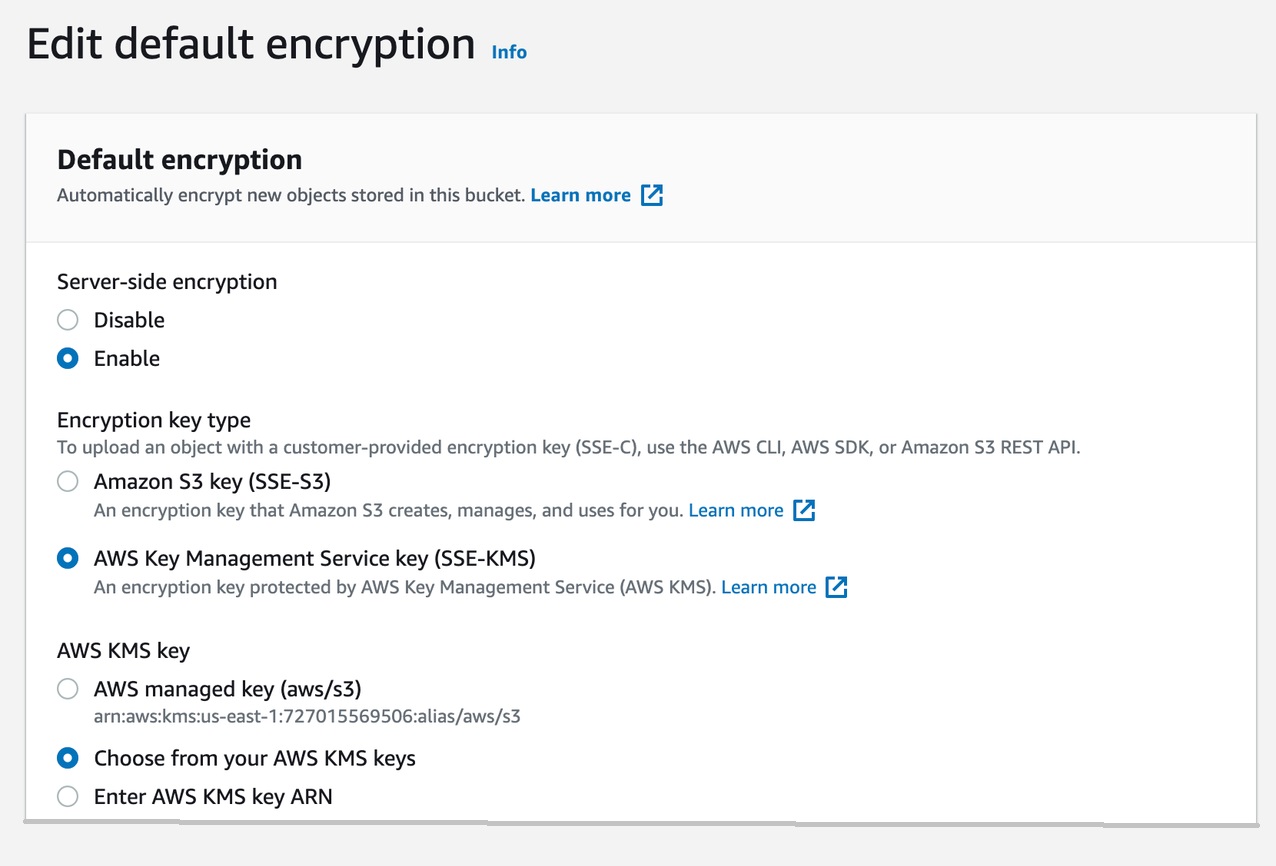
Then, enable server-side encryption for the S3 bucket that we’re using with CodeGuru Reviewer (Figure 9).
S3 bucket settings:

Figure 9: S3 bucket encryption settings
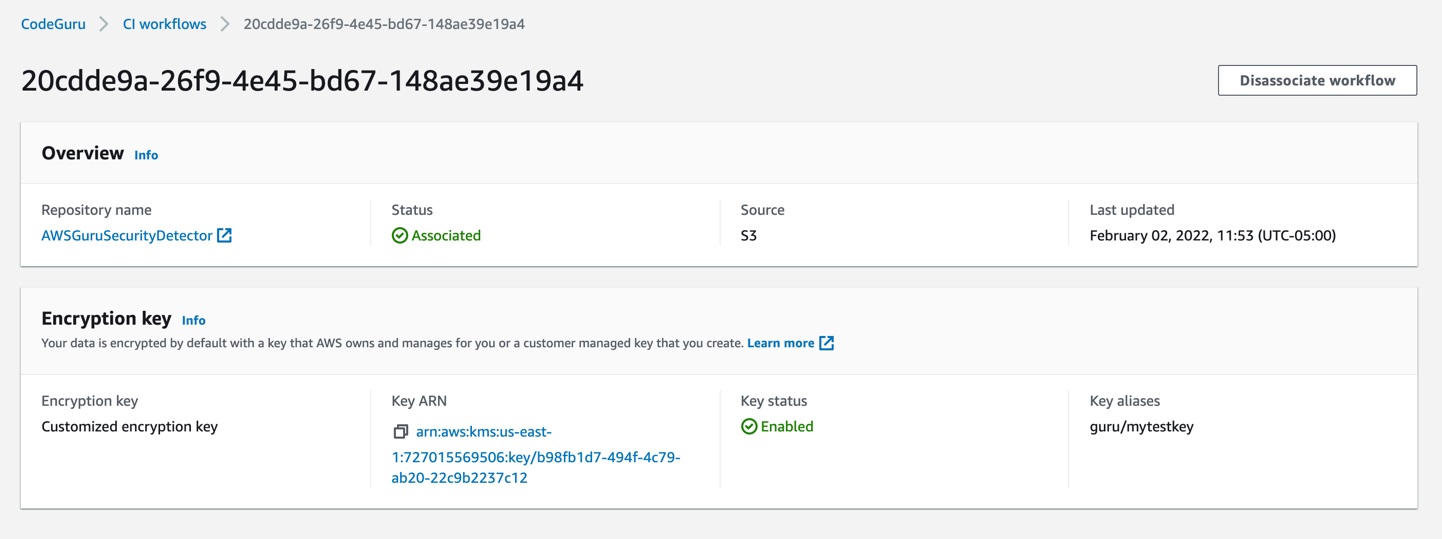
After we enable encryption on the bucket, we must delete all the CodeGuru repository associations that use this bucket, and then recreate them by analyzing the repositories while providing the key (as in the following example, Figure 10):

Figure 10: CodeGuru CI Workflow
Note that the first time you check out your repository, it will always trigger a full repository scan. Consider setting the -c option, as this will allow a commit range.
Cleaning Up
At this stage, you may choose to delete the resources created while following this blog, to avoid incurring any unwanted costs.
- Delete Amazon S3 bucket.
- Delete AWS KMS key.
- Delete the Jenkins installation, if not required further.
Conclusion
In this post, we outlined how you can integrate Amazon CodeGuru Reviewer CLI with the Jenkins open-source build automation tool to perform code analysis as part of your code build pipeline and act as a quality gate. We showed you how to create a Jenkins pipeline job and integrate the CodeGuru Reviewer CLI to detect issues in your Java and Python code, as well as access the recommendations for remediating these issues. We presented an example where you can stop the build upon finding critical violations. Furthermore, we discussed how you can specify a commit range to avoid a full repo scan, and how the S3 bucket used by CodeGuru Reviewer to store artifacts can be encrypted using customer managed keys.
The CodeGuru Reviewer CLI offers you a one-line command to scan any code on your machine and retrieve recommendations. You can run the CLI anywhere where you can run AWS commands. In other words, you can use the CLI to integrate CodeGuru Reviewer into your favourite CI tool, as a pre-commit hook, or anywhere else in your workflow. In turn, you can combine CodeGuru Reviewer with Dynamic Application Security Testing (DAST) and Software Composition Analysis (SCA) tools to achieve a hybrid application security testing method that helps you combine the inside-out and outside-in testing approaches, cross-reference results, and detect vulnerabilities that both exist and are exploitable.
Hopefully, you have found this post informative, and the proposed solution useful. If you need helping hands, then AWS Professional Services can help implement this solution in your enterprise, as well as introduce you to our AWS DevOps services and offerings.
About the Authors